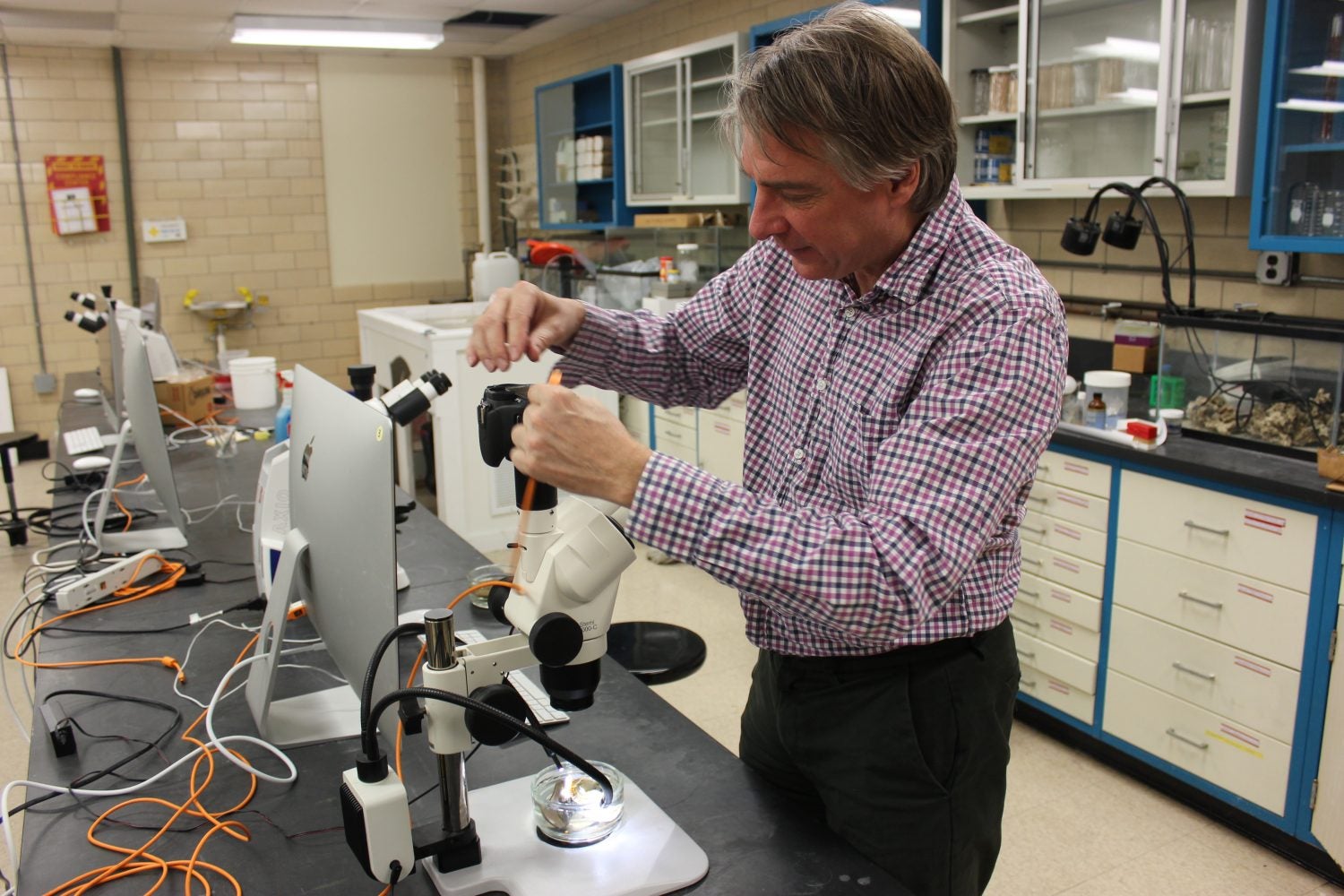

For the past 30 years, Dr. Joseph DeGiorgis has been face-to-face with the vast variety of marine organisms living in the waters of Narragansett Bay as a scuba diver for the Marine Biological Laboratory (MBL) in Woods Hole. Now, the Professor of Biology at Providence College is part of a collaborative effort among seven higher education institutions across Rhode Island to catalog all species of Narragansett Bay through a technique called ‘DNA barcoding.’
“We take a little piece of tissue from each organism and isolate their genomic DNA,” explains DeGiorgis. “It takes about 45 minutes, a very simple process.”
Through what is called a PCR reaction, scientists can amplify DNA strands of a particular species, which in turn provides a ‘molecular signature.’
“It’s a thumbprint for that species,” notes DeGiorgis.
The Providence College professor is joining a sizeable group of engineers, scientists, artists, and others on RI C-AIM (Rhode Island Consortium for Coastal Ecology—Assessment, Innovation, and Modeling), a five-year, $19 million initiative funded through the RI NSF EPSCoR program to develop a technological infrastructure for predicting and responding to the changing interactions between chemicals and lifeforms in Narragansett Bay.
One of RI C-AIM’s specific goals is to promote collaboration among scientists and artists to develop unique, visual platforms for presenting data and fostering broader understanding of scientific observations.
DeGiorgis has cameras ready to take high-resolution images of each marine species that will complement the barcoding initiative, and he has even created a special mount for the microscopes and cameras on-hand at the newly formed PC Aquatic Studio.
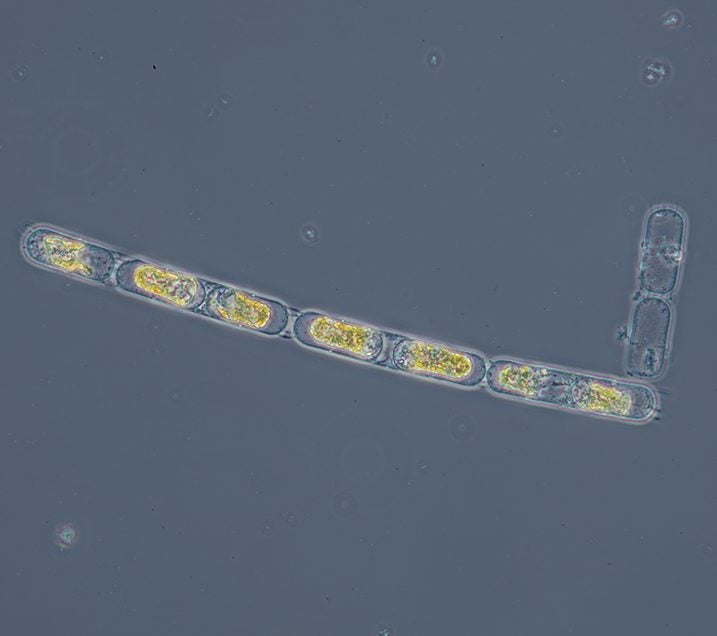
“I would like [the project] to include a pictorial library which will contain photographs that not only provide a scientific representation for each species, but also captures the aesthetic beauty of this marine life,” he says.
The Woods Hole resident is working side-by-side with RI C-AIM’s Neal Overstrom, director of the Edna Lawrence Nature Lab at the Rhode Island School of Design, as well as Oceanographer/Taxonomist Dr. Jan Rines at the University of Rhode Island, to match the barcoding results with known species.
“You really need those taxonomic expertsbecause just having a photograph of the critter and the DNA barcode doesn’t mean anything unless you accurately know what the species is,” emphasizes DeGiorgis. “Success with these organisms will require collaboration, and there is no real way to tell when you have every species. What if a jellyfish from Florida swims into the bay during the summer one year and then doesn’t show up again for a decade?”
But are the species of Narragansett Bay not already well-known? Yes and no, says DeGiorgis.
“One of my favorite papers estimates that the number of species on the planet is approximately 8.7 million, but only about 1.8 million currently have been identified taxonomically,” he says. “To me, it is so crazy to think that if you were doing DNA barcoding, there is a very good chance you will find novel species.”
“In New England, you have three world-class marine institutes—MBL, Woods Hole Oceanographic Institute, and the University of Rhode Island—piled on top of each other studying mostly the same critters, so Narragansett Bay could be the best studied body of water in the world. But there is still a possibility.”
To date, DeGiorgis’ team has collected DNA barcoding information on 60 organisms, but with thousands of species yet to be sampled, the goal is that the project can be sustained beyond C-AIM’s current five-year grant.
Cataloging the bay’s organisms, from the tiniest diatoms to the largest fish, will take time, but DeGiorgis notes that much of the process is simple and inexpensive, and thus involving students and citizen scientists in helping RI C-AIM researchers collect and code species is a real possibility.
“It is a very grassroots type of thing,” says DeGiorgis. “At the extreme ends of it, you can have top experts looking at the taxonomy and phylogenetic relationships of these species as well as junior high and high school kids and hobby scientists.”
“I think the project has all of this potential for outreach, getting local communities and schools involved at all levels. We would very much welcome others to participate.”
For more information about DeGiorgis’ work and RI C-AIM, visit https://web.uri.edu/rinsfepscor/ or contact DeGiorgis at jdegiorg@providence.edu.
Written by Shaun Kirby, RI C-AIM Communications & Outreach Coordinator

 RI NSF EPSCoR is supported in part by the U.S. National Science Foundation under EPSCoR Cooperative Agreements #OIA-2433276 and in part by the RI Commerce Corporation via the Science and Technology Advisory Committee [STAC]. Any opinions, findings, conclusions, or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the U.S. National Science Foundation, the RI Commerce Corporation, STAC, our partners or our collaborators.
RI NSF EPSCoR is supported in part by the U.S. National Science Foundation under EPSCoR Cooperative Agreements #OIA-2433276 and in part by the RI Commerce Corporation via the Science and Technology Advisory Committee [STAC]. Any opinions, findings, conclusions, or recommendations expressed in this material are those of the author(s) and do not necessarily reflect the views of the U.S. National Science Foundation, the RI Commerce Corporation, STAC, our partners or our collaborators.