Staphylococcus aureus (S. aureus) is a common bacterium found in the nasal cavity. It can cause severe infections and its MRSA strain is highly resistant against antimicrobial drugs.
A team of researchers at the University of Rhode Island has been studying the structure and dynamics of the S. aureus cell membrane. Their work can enable more effective drugs to be designed to counteract it.
Associate Professor Michael Greenfield, chemical engineering doctoral graduate Faramarz Joodaki, and Associate Professor Lenore Martin described the results of their research in the prestigious journal Langmuir.
Titled “Generation and Computational Characterization of a Complex Staphylococcus aureus Lipid Bilayer,” the article was published in August, with related images displayed on the cover.
Greenfield, who teaches chemical engineering at URI, summarized the article below.
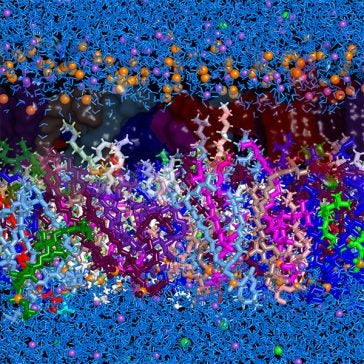
The cell membrane of S. aureus, like that of many organisms, is a bilayer of lipid molecules: what we colloquially call “fats.” It distinguishes “outside” from “inside” through two water-liking interfaces that surround a thin water-hating region.
In reality, the molecule types in a bilayer depend on factors such as the surrounding environment, the growth stage, and the intrinsic ability of an organism to synthesize or modify existing lipids. In simple representations of a lipid bilayer, this complexity is replaced by using a single type of lipid in experiments or in computer models.
Prior studies demonstrated that S. aureus changes its bilayer composition, leading to acquired antibiotic resistance and changes in membrane permeability.
The hypothesis of this work from URI is that important features of S. aureus lipid bilayer biophysics can emerge when some complexity of the real bilayer is incorporated into a model system, which ultimately could help to improve understanding of antibiotic resistance and to design better antimicrobial drugs.
For the first time, Joodaki et al. devised a composition of the S. aureus lipid bilayer that represents diverse experimental data on a computer. They used 19 different phospholipids and incorporated composition distinctions between the inside and outside of the bilayer. Such distinctions have been measured in experiments as one example of how S. aureus responds to external stress, such as the presence of antibiotics.
Next, the URI group ran large-scale molecular dynamics calculations to simulate how the bilayer evolved over 0.6 microseconds, or 0.6 millionths of a second. This is very short on the time scale of a person yet long compared to the internal time scales of a lipid bilayer. Averages of the motions and relaxations over this time form the basis of their interpretations of the S. aureus lipid bilayer.
Joodaki et al. reported several findings that could not be identified with a simpler bilayer of only one lipid type. The tails of the inner and outer halves of the bilayer overlapped, rather than forming a distinct boundary. Molecule branches near the end of each tail – a feature common in bacteria such as S. aureus though rare in most species – increased fluidity. This mechanism is distinct from ways that double bonds of unsaturated fats induce fluidity in mammalian lipid bilayers.
An important outcome of this work is its demonstration of how differences in bilayer composition exert differences in how a bilayer relaxes and moves, compared to simpler bilayers that only incorporate lipids that are not as common in bacteria.
Antibiotics and anti-microbials function through direct interactions and biochemical reactions with the lipid bilayer. Using a bilayer in model studies whose properties and responses are more typical of a real bacterium should provide more indicative results about how real bacteria can respond to new proposed therapies.
